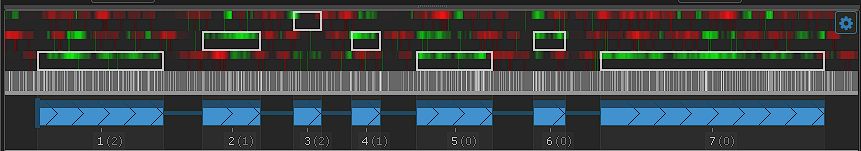
Feature update: Edit Gene Models
The latest addition to Persephone’s toolset is an interactive interface for editing gene models. When you encounter an issue in an existing gene model, you can now create your own corrected version of the gene structure and save it so others can review and use it.
We intentionally do not allow direct deletion or modification of existing genes, since those records may already be referenced in reports, analyses, or automated pipelines. Instead, alternative structures can be proposed in a dedicated track called Manual annotation, ensuring that community‑driven improvements are captured without altering the original data.
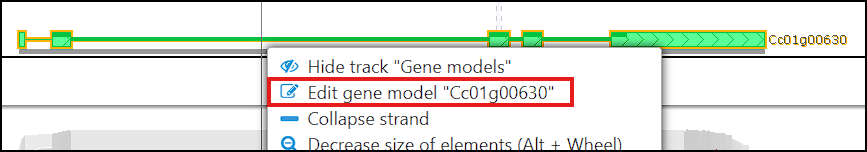
Right‑clicking a gene glyph in an annotation track and choosing “Edit gene model” opens (or creates, if needed) the Manual annotation track and brings up an editing form. This interface lets you adjust exon–intron boundaries directly, providing a controlled way to propose an updated gene structure.
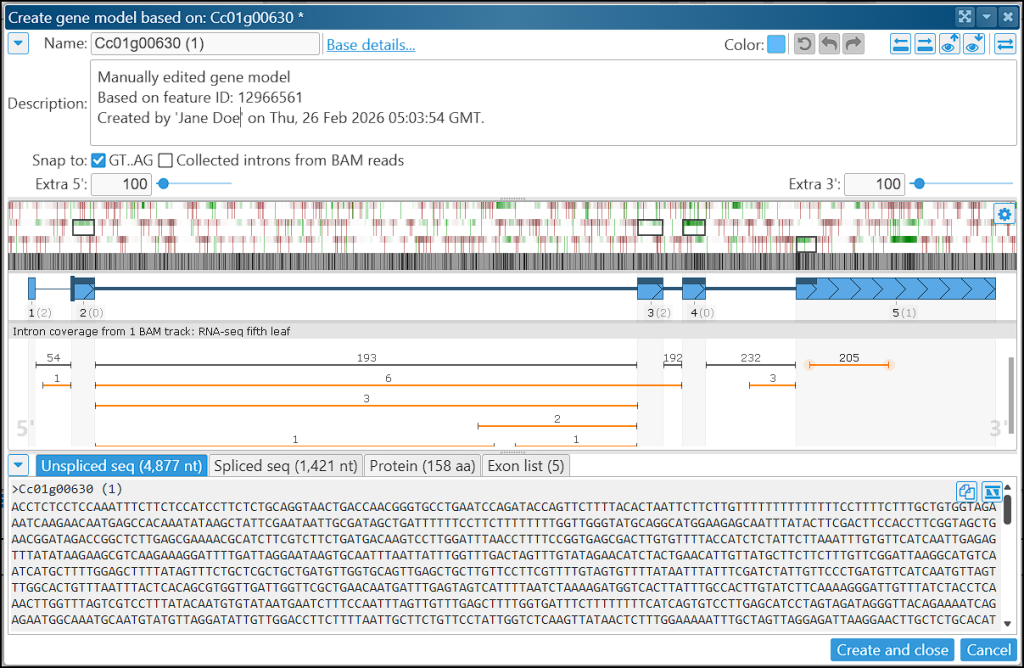
You may notice that the interface is quite busy, so it helps to keep unused panels collapsed. Each panel has a small triangle in its top‑left corner that toggles its collapsed state.
Above the gene model in the center of the form, you’ll see the sequence track showing coding potential across all three reading frames. Green highlights indicate regions and frames with a high likelihood of being protein‑coding. The coding potential is computed in real time from all models in the track and applied across the three frames in the current region. The values are based on the log‑odds ratio of hexamer frequencies in coding versus non‑coding sequences. Sensitivity and smoothing window size can be adjusted in the Settings popup.
The panel below the model displays intron junctions derived from RNA‑seq alignments, typically provided through a BAM/CRAM track. The numbers above each junction glyph represent the number of reads supporting that splice junction. Junctions that match the current gene model are shown in dark color; mismatching junctions are drawn in orange. If the intron flanking sequences are non‑canonical (not GT–AG), they are marked with orange (AT–AC) or red circles.
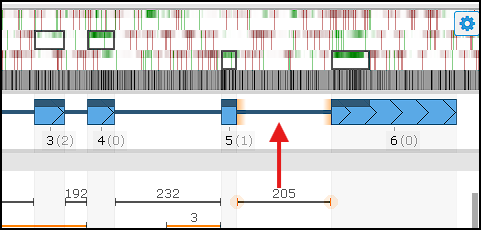
In the example above, you can see a strong coding‑potential signal within the 3′ UTR:
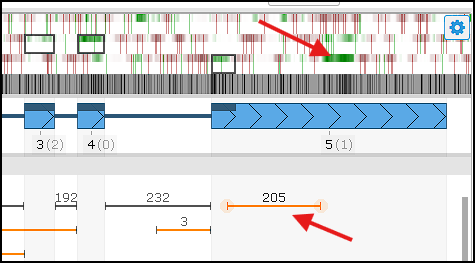
At the same time, there is strong support (205 reads) for an intron that extends the CDS and incorporates this high‑potential coding region. To update the gene model and add the intron, double‑click the junction glyph or choose the corresponding option from the right‑click menu. The revised model aligns more closely with the evidence and yields a longer predicted peptide.
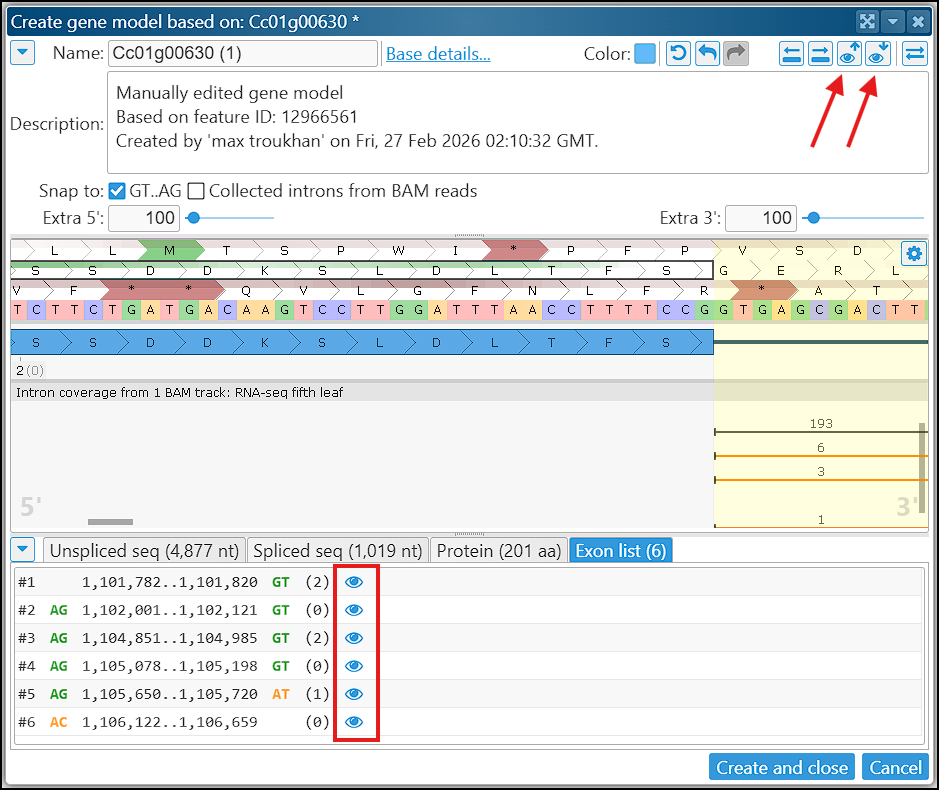
Editing gene models can be done either in this dedicated form or directly within the Manual annotation track. Exon and intron boundaries can be adjusted by dragging the edges of individual exons. Every step in modification causes the CDS to be recalculated automatically. If needed, you can drag the start codon position manually, which will determine the rest of the CDS and the stop codon. One of the tabs at the bottom of the form lists all exons, including their intron‑flanking dinucleotides, and provides “eyeball” icons for quick navigation to each exon. The gene views in the editing form and in the track can be synchronized in both directions – from the form to the track and vice versa.
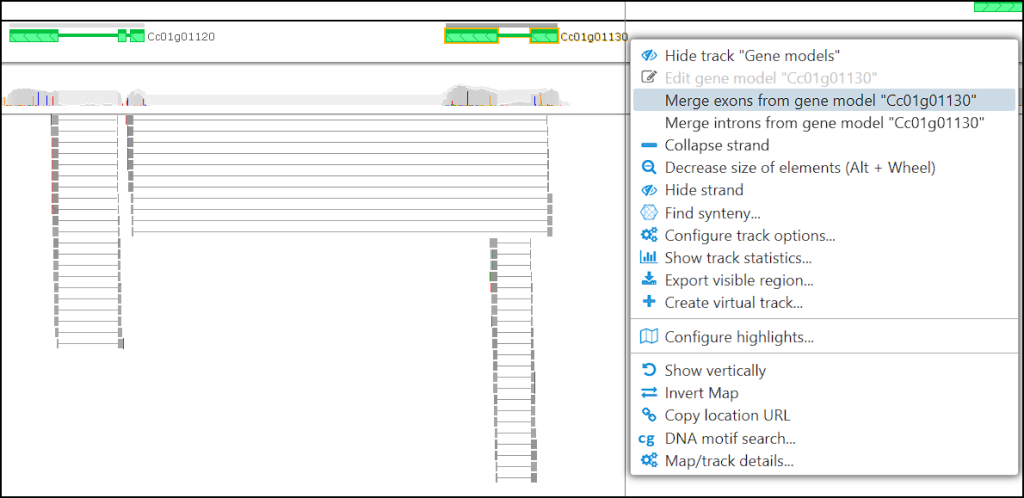
To join two genes, first, start editing one of the models and then right-click on the second model. A menu item will allow to “merge exons” or “merge introns”. This way, you can transfer exons from one model to another. If the genes do not overlap, by merging exons you can make a fusion of two or more genes.
If needed, you can also create a new model in an empty region. Highlight a segment of the sequence (Ctrl‑drag) and choose “Create gene model” from the right‑click menu. The program will generate a single‑exon gene spanning the highlighted interval and propose the longest open reading frame as its CDS. You can then introduce introns based on supporting evidence and save the completed model.
Please note that saving gene models is disabled in the free web application at https://web.persephonesoft.com. You can still explore the interface, experiment with the editing tools, and share any feedback you may have.
A page in the user guide describes this interface in detail.



