Please welcome: barley.persephonesoft.com
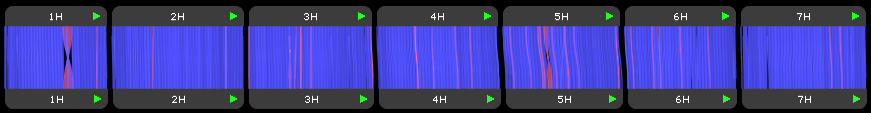
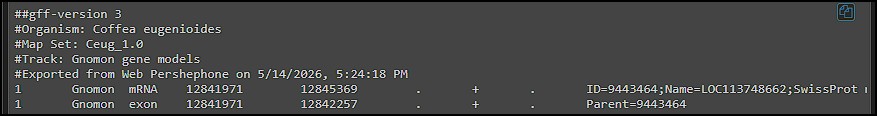
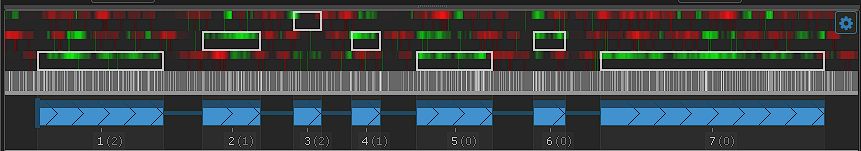
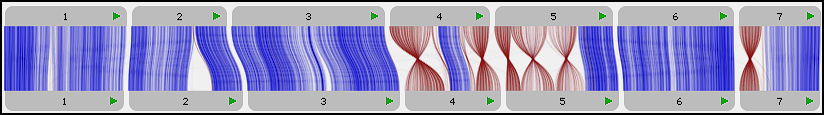
We are excited to announce a new, separate portal, specifically for barley pangenome data: https://barley.persephonesoft.com. It hosts 76 Hordeum vulgare assemblies described in the paper by Jayakodi, M., Lu, Q., Pidon, H., et al. Structural variation in the pangenome of...