Feature Update: Design SNP Assay (updated)
Updated: added batch processing
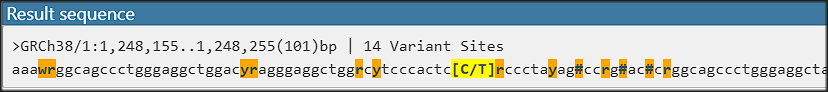
If you design an assay targeting a specific allele, it is important to know the genomic sequence neighborhood. A new addition to the Local variants interface will allow users to export flanking sequences around the selected variant. The extracted sequence can be formatted differently and typically will use IUPAC codes for variant sites in the flanking sequence while splitting the alleles of the ‘anchor’:

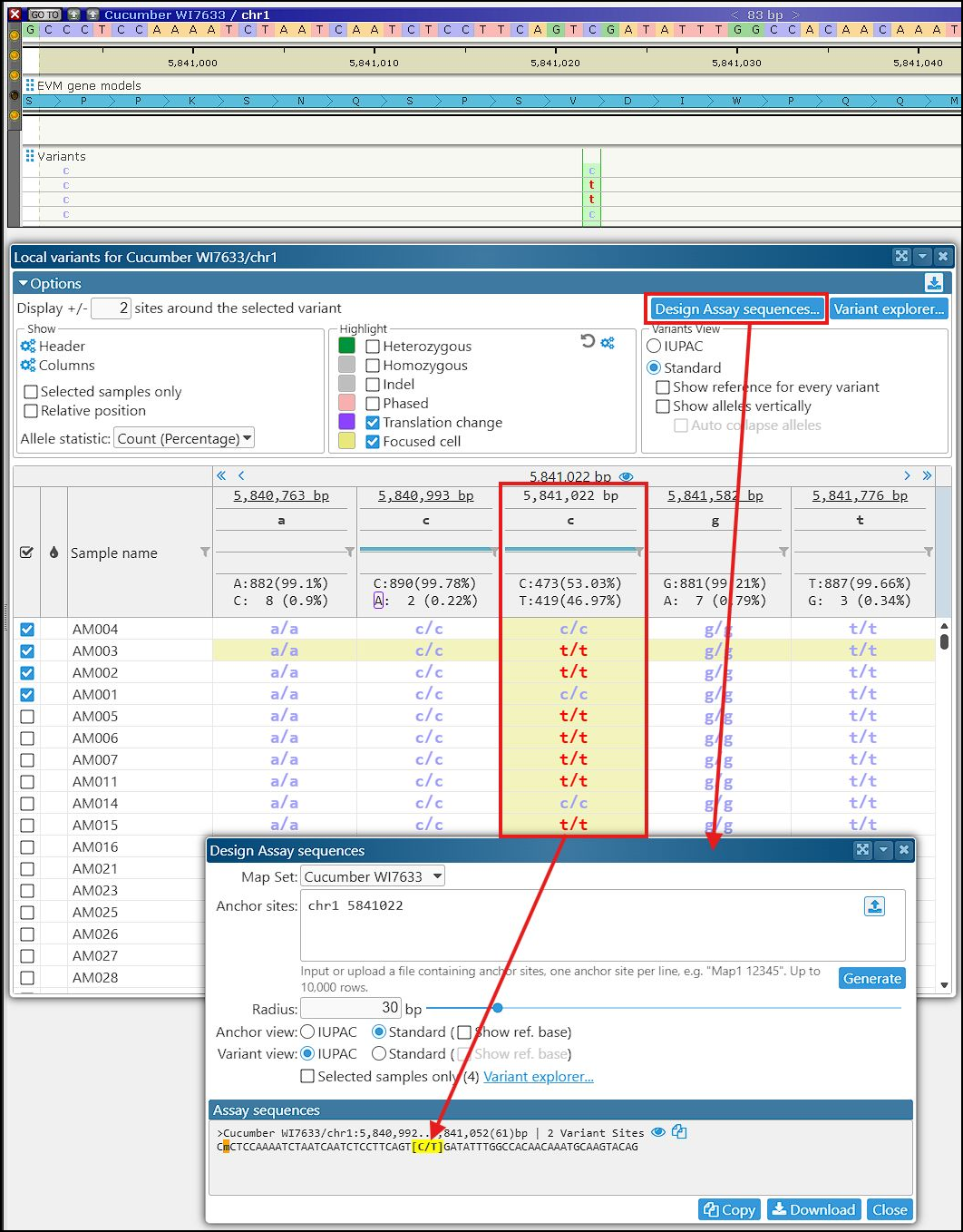
Clicking the eye icon in the sequence header will display the assay region around the anchor site on the map.
The interface for designing SNP assays can also be accessed from the Tools menu and from Variant Explorer. Note that the text field Anchor sites can be used for batch processing. If you have a list of anchor sites with MapName and Coordinate, you can paste or upload it to the form. This will allow you to generate up to 10,000 assay sequences.



